-

How to Analyze RNAseq Data for Absolute Beginners Part 15-2: Mastering UMI-Based miRNA-Seq Analysis
Understanding UMI-Based miRNA Sequencing MicroRNAs (miRNAs) serve as crucial regulators in gene expression, making their accurate quantification essential for understanding disease mechanisms and biological processes. While traditional miRNA sequencing has proven valuable, the integration of Unique Molecular Identifiers (UMIs) represents a significant advancement in achieving precise miRNA measurements. This tutorial will guide you through the
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 15: A Complete Guide to miRNA-seq Analysis
Understanding the World of microRNAs The fascinating world of microRNAs (miRNAs) represents one of molecular biology’s most elegant regulatory systems. These tiny RNA molecules, spanning just 20-24 nucleotides, function as precise genetic regulators by binding to messenger RNAs (mRNAs) and fine-tuning their expression. Since their serendipitous discovery in the early 1990s, miRNAs have revolutionized our
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 14: Mastering Small RNA-Seq Analysis
The Hidden World of Small RNAs: More Than Just Tiny Molecules Small RNAs are fascinating molecules that challenge the traditional “DNA to RNA to protein” dogma of molecular biology. Despite being just 20-30 nucleotides in length – barely a fraction of a typical messenger RNA – these molecules orchestrate complex biological processes with remarkable precision.
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 13: Circular RNAseq Analysis
Understanding the Biology of Circular RNAs The Nature of Circular RNAs Circular RNAs (circRNAs) represent one of molecular biology’s most fascinating discoveries. Unlike the linear RNA molecules that dominated our understanding of gene expression for decades, circRNAs form continuous loops through a unique process called back-splicing. In this process, a downstream 5′ splice site connects
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 12: A Step-By-Step Guide for Submitting Your NGS Data to NCBI GEO
In the world of genomics research, sharing your sequencing data isn’t just a box to check for publication – it’s a fundamental part of advancing scientific knowledge. As researchers, we spend countless hours generating and analyzing Next-Generation Sequencing (NGS) data, and making this data accessible to the broader scientific community ensures our work can have
//
-
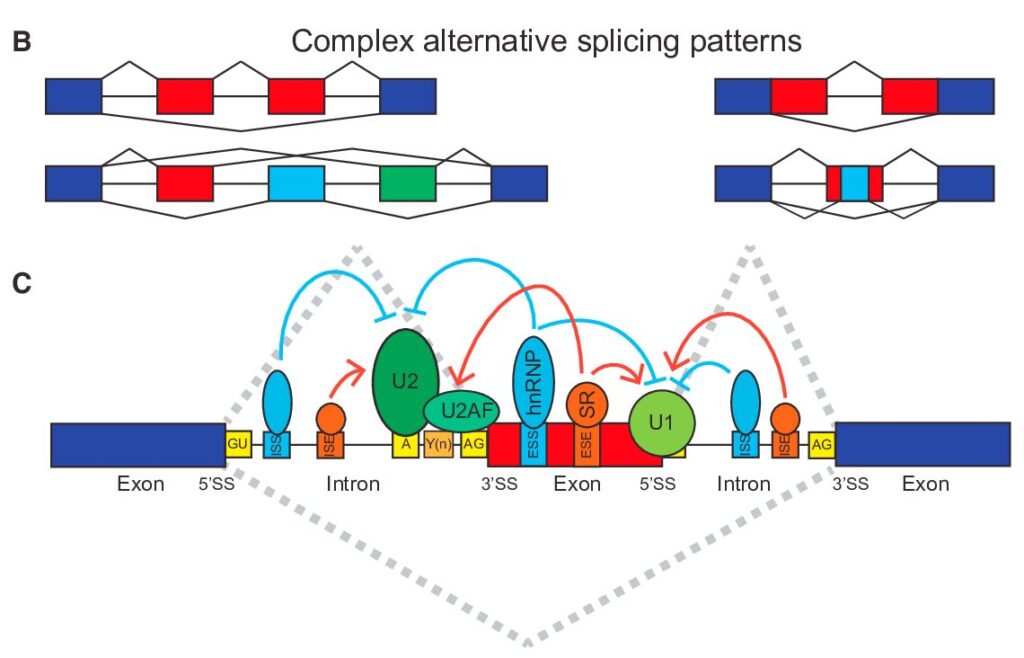
How to Analyze RNAseq Data for Absolute Beginners Part 11: Mastering Transcript-Level Alternative Splicing Analysis
From Gene-Level to Transcript-Level Analysis In our previous exploration of gene-level splicing analysis, we laid the groundwork for understanding how alternative splicing shapes gene expression. Now, we’re taking a deeper dive into the fascinating world of transcript-level analysis, where we can uncover the intricate details of how genes produce different protein variants through alternative splicing.
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 10: Isoform Analysis
If you’ve followed our previous tutorials on RNA-seq analysis, you’re already familiar with gene-level analysis. But genes are far more complex than simple on/off switches – they can produce multiple versions of RNA transcripts through fascinating processes like alternative splicing. These different versions, called isoforms, allow a single gene to create multiple protein products, dramatically
//
-
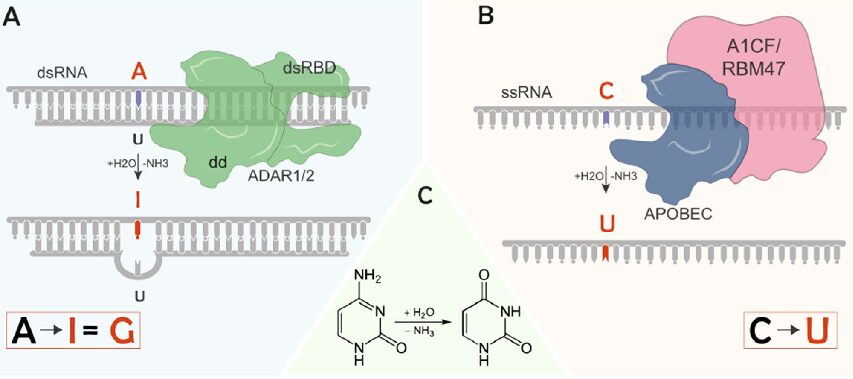
How to Analyze RNAseq Data for Absolute Beginners Part 9: RNA Editing Analysis
RNA editing represents one of the most fascinating mechanisms in molecular biology – a process that introduces targeted changes to RNA sequences after transcription, creating a dynamic transcriptome that diverges from the genome’s blueprint. Through this tutorial, you’ll learn how to analyze RNA editing events from RNA-seq data, uncovering these crucial modifications that fine-tune gene
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 8: Alternative Splicing Analysis
Introduction The human genome harbors an elegant solution to the challenge of biological complexity. Through alternative splicing (AS), a single gene can orchestrate the production of multiple protein variants, much like a composer creating different melodies from the same set of musical notes. This remarkable mechanism serves as nature’s way of expanding our proteome’s diversity,
//
-

How to Analyze RNAseq Data for Absolute Beginners Part 7: Unlocking Cell-Type Resolution from Bulk RNA-seq Data With Deconvolution Analysis
Understanding Bulk RNA-seq Deconvolution Analysis: Unraveling Cellular Complexity In the ever-evolving landscape of molecular biology, bulk RNA sequencing has emerged as a fundamental technology for understanding gene expression patterns. However, like peering through a frosted window, bulk RNA-seq provides only an averaged view of the complex cellular world within our tissues. This challenge has given
//
Search
Subscribe
Categories
- bulk RNA-seq (27)
- chromatin accessibility (14)
- Database (4)
- Epigenetics (14)
- Genomics (10)
- HPC (5)
- Metagenomics (1)
- Quick Tips (1)
- RNA-seq (15)
- Scientific Programming (5)
- Single Cell Sequencing (15)
- Transcriptomics (28)
Recent Posts
- How to Analyze Single-Cell RNA-seq Data — Complete Beginner’s Guide Part 13: RNA Velocity Analysis with scVelo
- How to Analyze Single-Cell RNA-seq Data – Complete Beginner’s Guide Part 12: Build Gene Co-expression Networks Using hdWGCNA
- How to Analyze Single-Cell RNA-seq Data — Complete Beginner’s Guide Part 11: Copy Number Variation Analysis Using CopyKAT
- No More Command-Line Only: Run Jupyter Lab, RStudio, and VS Code Interactively in Your Browser on Any HPC Cluster with Pixi
Tags
Alternative Splicing Analysis ATAC-seq BAM ChIP-seq chromatin accessibility CNV DESeq2 Differential Expression edgeR FASTQ GATK Mutect2 gene expression heatmap HOMER HPC Isoform limma MACS2 MAF miRNA miRNA-seq MSigDB Normalization peak calling RNA-seq SLURM somatic mutations Transcript VCF whole genome sequencing



